Dr. Yaron Carmi
Department of Pathology

Research
Dr. Yaron Carmi
Department of Pathology


Research
Prof. Arieh S. Solomon, M.D., Ph.D.
Goldschleger Eye Research Institute
Department of Ophthalmology

The eye presents many challenges for research regarding unsolved conditions such as retinal and optic nerve assaults, damage to eye by surrounding conditions of work and every day activity.
The following specific subjects are studied:
| 2012-2015 | European Union FP7 |
Research
Dr. Inna Slutsky, Ph.D.
Department of Physiology and Pharmacology

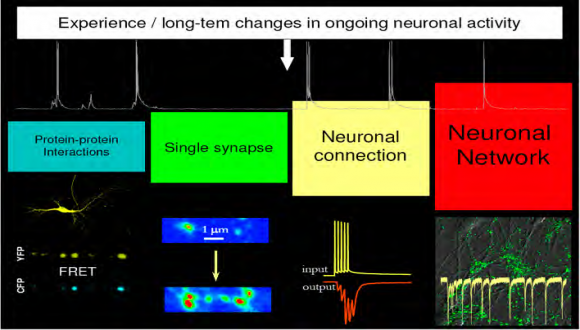
The research in the laboratory is focused on understanding the basic mechanisms underlying synaptic function and primary mechanisms initiating synaptic dysfunction at very early stages of Alzheimer’s Disease. To achieve this goal, we developed an integrated system that enables simultaneous real-time visualization of structural reorganization in spatially-restricted signaling complexes and functional modifications of single synapses in brain circuits.
Utilizing FRET spectroscopy, high-resolution optical imaging, electrophysiology, molecular biology, and biochemistry we explore experience-dependent mechanisms regulating the number and plasticity of hippocampal synapses under physiological and pathological conditions.
| 2011-2016 | Evolution of Alzheimer’s Disease: From Dynamics of Single Synapses to Memory Loss, European Research Council Starting Grant. |
Research
Prof. Moshe Rehavi, Ph.D.
Department of Physiology and Pharmacology

Main projects in the lab include:
Research
Prof. Chaim G. (Chagi) Pick, Ph.D.
Department of Anatomy and Anthropology

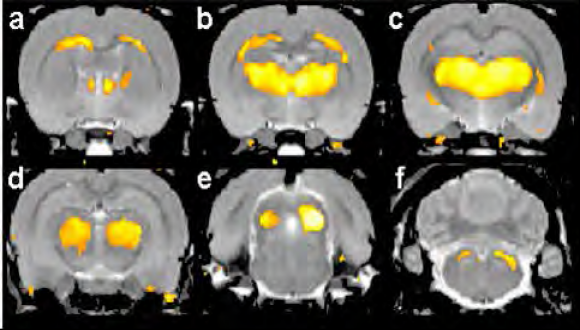
My group has a long history in mTBI research, not only in characterizing behavioral and biochemical sequelae of blunt head trauma, but also in developing preclinical models of mTBI of translational relevance to support the development of new treatment strategies and drugs. In order to look for answers regarding the blast induced traumatic brain injury, we have developed a blast injury model for mice that resembles, as much as possible, the conditions on the battlefield or at a terror-attack site. As such, the outcomes of the “real-life-like” exposure to the blast in our model may vary from severe to mild brain injury under controlled conditions for each mouse.
Research
Dr. Eran Perlson, Ph.D.
Department of Physiology and Pharmacology

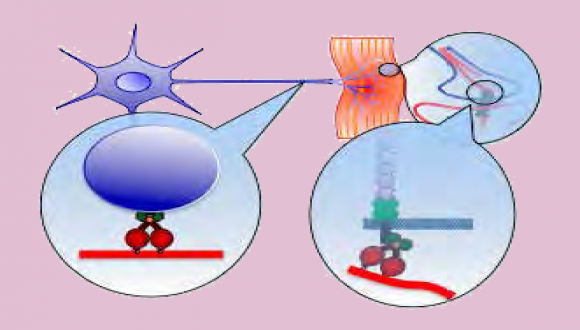
The lab is a new multi-disciplinary molecular and cellular neurobiology lab. The lab uses state-ofthe- art single molecule live imaging techniques on neuronal cultures, as well as biochemistry, cell biology and biophysics approaches on mouse model systems to study the role of axonal transport in neurodegenerative diseases, with an initial focus on ALS.
Neuronal survival and proper function depends on cell-cell communication mediated by ligandreceptor mechanisms. During neurodegenerative diseases such as Amyotrophic Lateral Sclerosis (ALS), there is considerable synapse/neuromuscular junction (NMJ) disruption and neuronal cell death. It is non-autonomous processes involve interactions between the neurons to its diverse extracellular microenvironments. The molecular basis for this neuronal dysfunction and death is still poorly understood. One possible reason is alterations in the nature, directed movement and spatial localization of vital extra and intracellular signals.
The long-term research goal of the lab is to understand the vital molecular communications mechanisms between the neurons and its environment. More specifically, we seek to understand the role that retrograde signaling plays in (1) neuronal survival and (2) synapse stability. We believe that our research will generate novel insights into neurodegenerative mechanisms and ultimately, provide a molecular basis for new drugs as well as delivery methods to treat a range of neurodegenerative diseases.
| 2011-2015 | ISF (Israel Science Foundation), The Dual Role of Dynein in GDNF Signaling |
| 2011-2015 | Marie Curie International Reintegration Grants (IRG), Retrogade Signaling. |
| 2013-2016 | Small Molecule Screen for Neuromuscular Junction Maintenance, Rosetrees Trust |
| 2013-2016 | E-Rare-2, European Research Projects on Rare Diseases driven by Young Investigators. Project Coordinator. The Molecular Basis of Neurodegeneration and Muscle Atrophy in ALS. (Co-PIs: Roded Sharan, TAU; Edgar Gomes, U of Paris; Marcus Kruger, Max Planck; Del Bene Fillippo, Ins Curie; Alberto Rodendo, 12th Oct Uni Hospital Madrid) |
| 2013-2018 | Molecular Communication Mechanism of Motor Neuron Survival and Synapse Maintenance, European Research Council (ERC) Starter Grant |
Research
Dr. Moshe Parnas, Ph.D.
Department of Physiology and Pharmacology

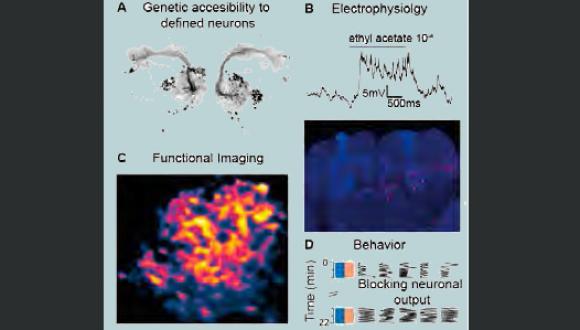
We are exploring the various mechanisms by which neural circuits encode information and support behavior, learning and memory. In addition, we are studying how the connectivity and activity of such circuits and neural networks are affected by molecular mechanisms underlying brain disorders. We use a multidisciplinary approach, with the Drosophila olfaction system as our model system. Our studies incorporate in vivo whole cell patch recordings, in vivo functional imaging, behavior experiments, molecular biology, mathematical modelling and genetics.
Projects in the lab include:
Research
Prof. Illana Gozes, Ph.D.
Department of Human Molecular Genetics and Biochemisty

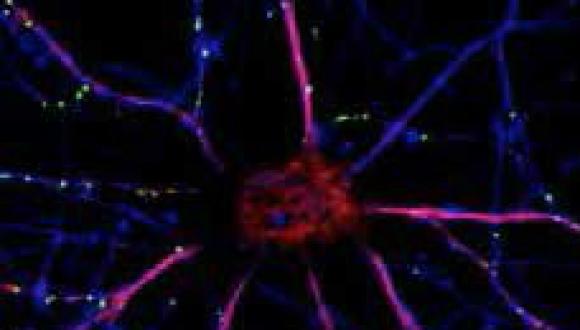
Our research is characterized by a multi-level approach to the study of brain function, behavior, memory and drug discovery, from molecules to cures. Targeting autism, schizophrenia as well as Alzheimer’s disease and related neurodegeneration and utilizing a multidisciplinary approach, our group investigates different aspects of neuronal plasticity and nerve cell protection, at the molecular, cellular and system level.
A major focus in the laboratory is on nerve structure and transport mechanisms. We have discovered novel families of proteins associated with cross talk among nerve cells and their support cells, including activity-dependent neurotrophic factor (ADNF) and activity-dependent neuroprotetive proteins (ADNPs, with ADNP being a major gene mutated in autism). Small ADNF and ADNP derivatives are in clinical development. The lead compound, davunetide is planned for an advanced Phase II clinical trial with the biotech industry.
Davunetide has previously shown efficacy in several Phase II clinical trials (i.e. in patients suffering from mild cognitive impairment, preceding Alzheimer’s disease and in schizophrenia patients, protecting activities of daily living).
Research
Prof. Hagit Eldar-Finkelman, Ph.D.
Department of Human Molecular Genetics and Biochemistry

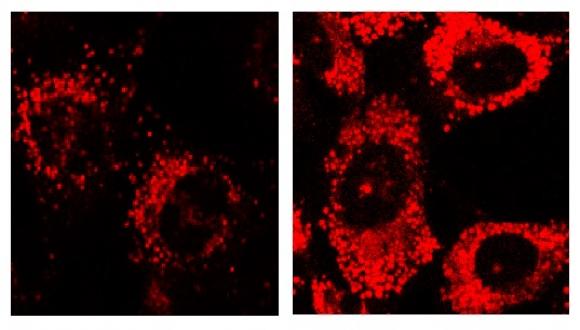
Our research is focused on the molecular mechanisms regulating the protein kinase GSK-3 and their implications in human disease. GSK-3 is a central player in diabetes, neurodegenerative and psychiatric disorders, and recently emerged as a promising drug discovery target. We propose that inhibition of GSK-3 should produce therapeutic benefits in treating these disorders. We develop selective substrate competitive GSK-3 inhibitors and evaluate their efficacy and therapeutic effects in relevant in vitro and in vivo systems. So far we could show that our leading compound inhibitors had therapeutic efficacy in CNS disorders models for Alzheimer’s disease, mood disorders, and multiple sclerosis.
In recent work we identified the lysosome as a GSK-3 target. This implicated GSK-3 as a key player in protein degradation pathways, particularly autophagy ad endocytosis. Research methods combine cell biology, molecular biology and biochemistry disciplines together with bioinformatics and computational biology.